Searching Pubmed using MeSH terms.
We saw in a previous post how to do a Pubmed search using the simplest system, which is to enter natural language text in the simple search box and press the “Search” button.
This method is quite easy and even works quite well when we are looking for something about very rare diseases but, in general, it will give us a very sensitive and unspecific results list, which in this context means that we will get a large number of articles, but many of them will have little to do with what we are looking for.
In these cases we will have to use some tool to make the result more specific: fewer articles and more related to the problem that originates the search. One of the ways is to perform an advanced search instead of a simple search, but for this we will have to use the browser’s own jargon, the so-called thematic descriptors of controlled language.
Some previous definitions
A descriptor is a term used to construct indexes, also called thesauri. Instead of using the words of the natural language, they are selected or grouped under specific terms, which are to serve as a key in the index of the search engine database.
The thesaurus, formed by the set of descriptors, is specific to each search engine, although many terms may be common. In the case of Pubmed the descriptors are known as MeSH terms, which are the initials of Medical Subject Headings.
This thesaurus or list of terms with controlled vocabulary has also been developed by the National Library of Medicine and constitutes another database with more than 30,000 terms that are updated annually.
Within the National Library there are a number of people whose mission is to analyze the new articles that are incorporated into the Medline database and assign them the descriptors that best fit their content. Thus, when we search using a particular descriptor, we will find the articles that are indexed with this descriptor.
Searching Pubmed using MeSH terms
But the thing of the descriptors is a little more complicated than it may seem, since they are grouped in hierarchies (MeSH Tree Structures), being able to a same descriptor to belong to several hierarchies, in addition to having subheadings, of such form that we can search using the general MeSH term or further narrow the search using one of its subheaders.
Reading all this we may want to forget the search using the thesaurus, but we cannot afford that lapse: the search using the MeSH database is the most effective and accurate, since the language has been controlled to eliminate inaccuracies and synonyms of natural language.
Also, the thing is not so complicated when we get to work with it. Let’s see it with the example we use to display the simple search. We want to compare the efficacy of amoxicillin and cefaclor on the duration of otitis media in infants. After elaborating the structured clinical question we obtain our five terms of search, in natural language: otitis, infants, amoxicillin, cefaclor and prognosis.
You can now go to the Pubmed home page. Below the area where the simple search window is, we saw that there are four columns with different information. We look at the one on the right, “Explore”, and we click on the first of the options, “MeSH Database”, with which we access the home page of the descriptor database (which we can see in the first figure ).
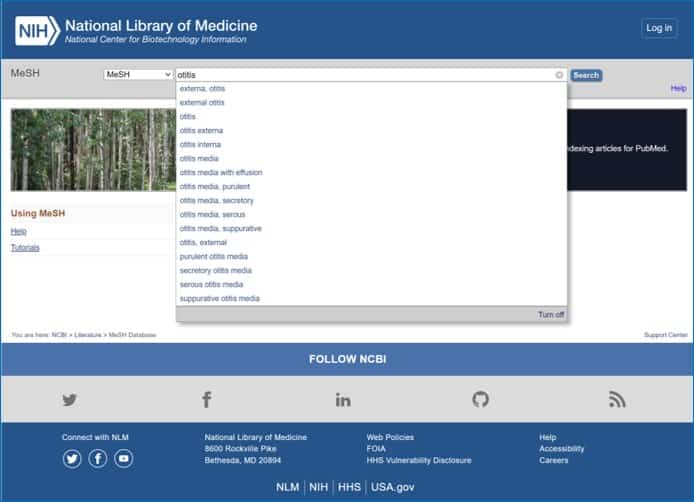
If we write otitis in the search window we see that Pubmed lends us a hand by displaying a list of terms that look like what we are writing. One of them is otitis media, which is what we are interested in, so we select it and Pubmed takes us to the next page where there are several options to choose from, as you can see in the attached figure.
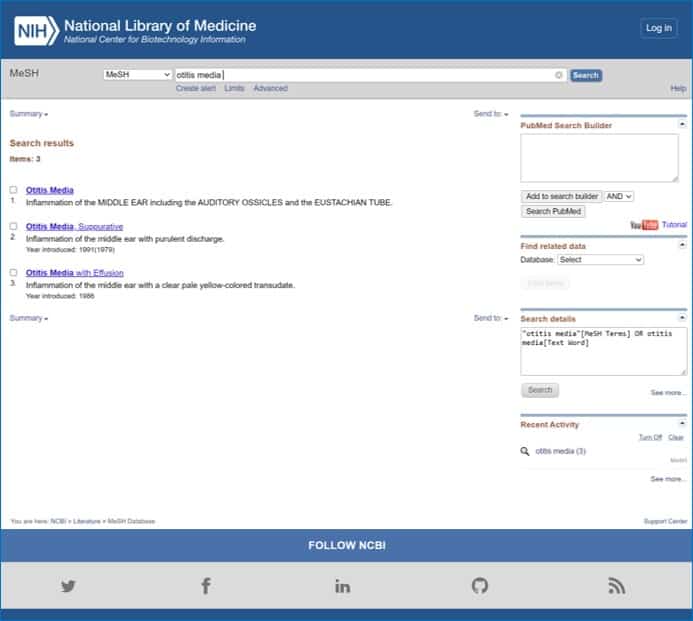
At the moment I do the search there are three options: “Otitis Media”, “Otitis Media, Suppurative” and “Otitis Media with Effusion”. Notice that Pubmed defines each term, so that we understand well what it means with each term. These are the three MeSH terms that fit what we asked for, but we have to choose one.
The simplest thing we can do from this window is to check the selection box to the left of the term that interests us and click the button on the right side of the screen that says “Add to search builder”.
If we do this, Pubmed begins to construct the search string starting with the chosen term. If we do this with the first term in the list you will see that the text “Otitis Media” [Mesh] appears in the text box “Pubmed Search Builder” , on the top right of the screen.
But remember that we have said that the MeSH terms have subheaders. To get them, instead of marking the selection box of the term “Otitis Media”, we click on the term, opening the window with the subheadings, as you can see in the third figure.
The hierarchical tree of MeSH terms
Each of the terms with their selection box on the left corresponds to a subheading of the descriptor “Otitis Media”. For example, if we were interested in doing a search directed to the cost of the treatment, we could mark the subheading “economics” and then press the button to add to the search. The text that would appear in the text box of the search string would be “Otitis Media / economics” [Mesh] and the search result would be a bit more specific.
Before leaving the MeSH term window let’s look at a few details. In addition to the subheadings, which can be more or less numerous, the bottom of the page shows the hierarchy of the descriptor (MeSH Tree Structure).
Our descriptor is in bold, so we can see which terms it depends on and which ones depend on it. In some cases we may be more interested in using a higher term for the search, so we will have to click on it to go to its own window. If we do this, in general, the search will be more sensitive and less specific (more empty vessels).
We can also click on a term that is below the hierarchy, making the search more specific and decreasing the number of results.
And it does not end here. If we select a MeSH term for the search, it includes the terms that are below in the hierarchy. For example, if we select the descriptor “Otitis Media”, Pubmed will include in the search all that hang from it (mastoidits, otitis with effusion, suppurative otitis and petrositis, which may not interest us at all). This can be avoided by checking the box that says “Do not include MeSH terms found under this term in the MeSH hierarchy”.
An example to finish
Well, I think we’re going to end up with this example, if there is still someone who is still reading at this point. Let’s say we chose the simplest way: let’s go to “Otitis Media” and add it to the search. Next we write the second search term in the search window of the database: infants. We get 41 possibilities, select the first (“Infant”) and add it to the search. We do the same with “Amoxicillin”, “Cefaclor” and “Prognosis”.
When we have added all of them to the search string (note that the default boolean operator is AND, but we can change it), the search string is as follows: (“(Otitis Media [Mesh]) AND” Infant ” Mesh]) AND “Amoxicillin” [Mesh]) AND “Cefaclor” [Mesh]) AND “Prognosis” [Mesh] (see attached figure).
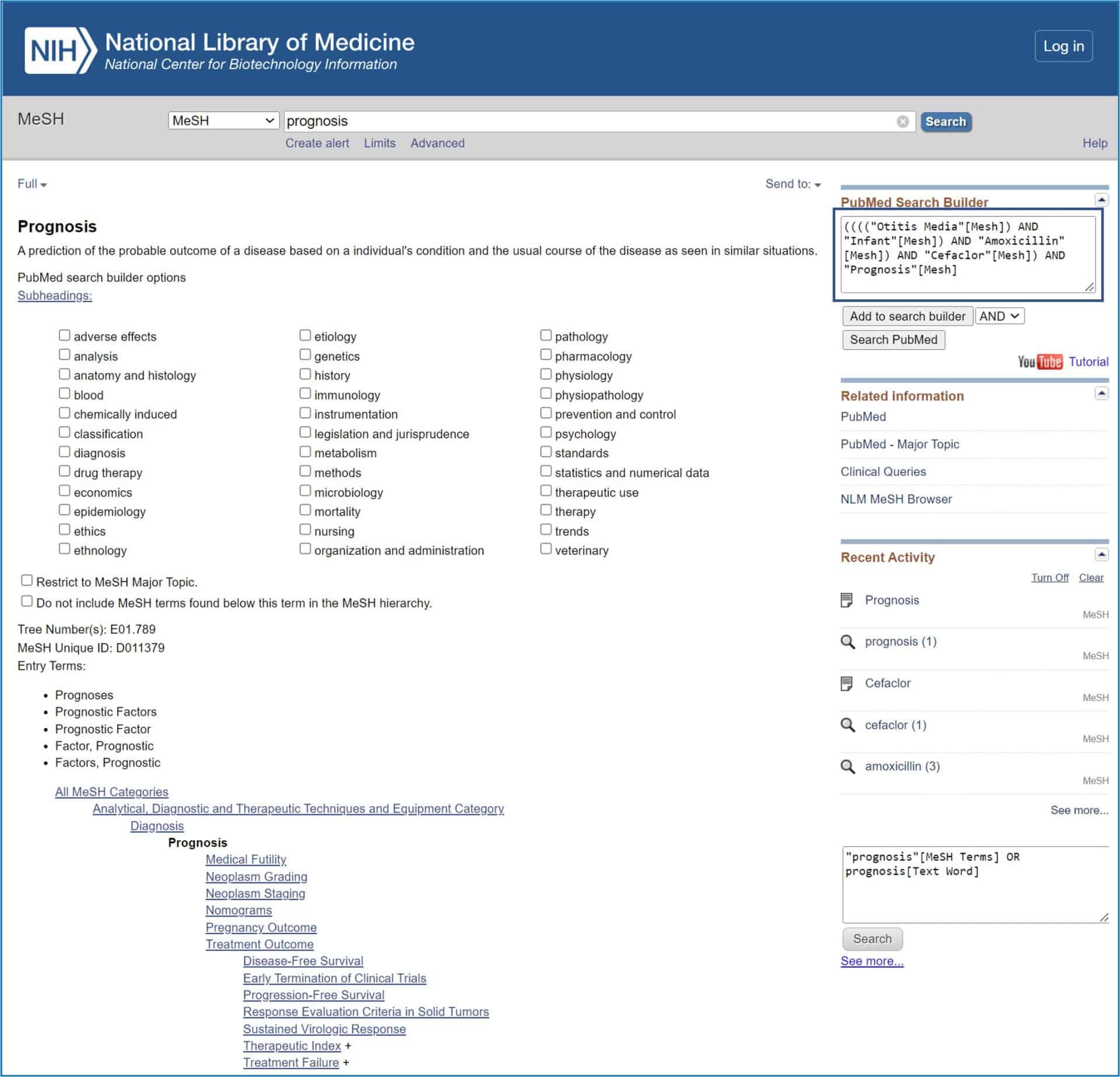
Finally, click the “Search PubMed” button and get the search result, which in this case is a bit more restricted than we obtained with natural language (this is usually the case).
If we wanted to remove the articles about the treatment with clavulanic acid, as we did in the example with the simple search, we could add the term clavulanate as we add the other terms, but changing the boolean operator AND by the NOT operator. But there is another way that is even simpler.
If you notice, when Pubmed gives us the list of results, in the search window of Pubmed is written the search string that has been used and we can add or remove terms from this string, using MeSH or natural language terms, which we prefer. So, in our example, to the text string we would add NOT clavulanate in the search box and we would click on the “Search” button again.
We’re leaving…
And here we are going to leave it for today. Just saying that there are other ways to use MeSH terms, using the advanced search form, and we can further narrow the results using some resources, like the Clinical Queries or using limits. But that is another story…